Powder: -20°C for 3 years | In solvent: -80°C for 1 year
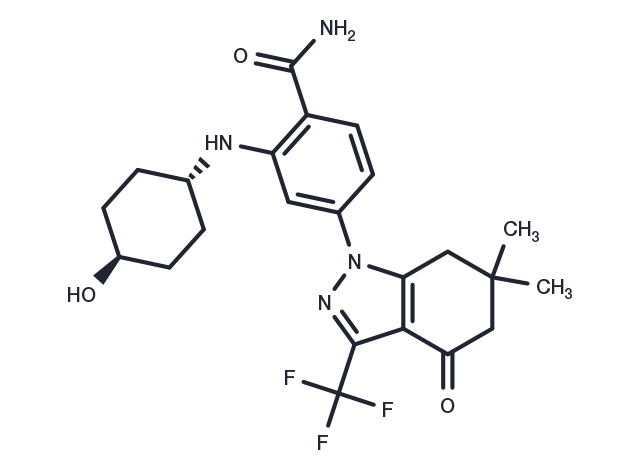
SNX2112 (PF 04928473) 是一种具有口服活性的 Hsp90 抑制剂,Kd 为 16 nM。
| 规格 | 价格/CNY | 货期 | 数量 | |
|---|---|---|---|---|
| 1 mg | ¥ 668 | 现货 | ||
| 2 mg | ¥ 981 | 现货 | ||
| 5 mg | ¥ 1,630 | 现货 | ||
| 10 mg | ¥ 2,250 | 现货 | ||
| 25 mg | ¥ 3,790 | 现货 | ||
| 50 mg | ¥ 5,460 | 现货 | ||
| 100 mg | ¥ 7,670 | 现货 | ||
| 500 mg | ¥ 15,300 | 现货 | ||
| 1 mL * 10 mM (in DMSO) | ¥ 1,810 | 现货 | ||
| 产品描述 | SNX2112 (PF 04928473) is an orally active Hsp90 inhibitor, with a Kd of 16 nM. |
| 靶点活性 | HSP90 β:30 nM(Ka), HSP90 α:30 nM(Ka) |
| 体外活性 | Treatment of BT-474 cells with 1 μM SNX-2112 results in down-regulation of HER2 expression within 3 to 6 hours of drug exposure with near-complete loss of HER2 expression by 10 hours. Treatment with SNX-2112 also results in a decline in total Akt expression. SNX-2112 inhibits cell proliferation with IC50 values ranging from 10 to 50 nM, in BT474, SKBR-3, SKOV-3, MDA-468, MCF-7 and H1650 cancer cells. And these antiproliferative effects are associated with hypophosphorylation of Rb, arrest of G1 and modest levels of apotosis. [1] SNX-2112 competitively binds to the N-terminal adenosine triphosphate binding site of Hsp90. SNX-2112 induces apoptosis via caspase-8, -9, -3, and poly (ADPribose) polymerase cleavage. SNX-2112 inhibits cytokine-inducedAkt and extracellular signal-related kinase (ERK) activation and also overcomes the growth advantages conferred by interleukin-6, insulin-like growth factor-1, and bone marrow stromal cells. SNX-2112 inhibits tube formation by human umbilical vein endothelial cells via abrogation of eNOS/Akt pathway and markedly inhibits osteoclast formation via down-regulation of ERK/c-fos and PU.1. [2] Cell lines (eight cell lines from osteosarcoma, neuroblastoma, hepatoblastoma, and ymphoma) studied demonstrates sensitivity to SNX-2112 with IC50 values ranging from 10-100?nM. A higher dose (70?nM) exhibits a more prolonged inhibition and larger sub-G1 accumulation. Observed levels of Akt1 and C-Raf are markedly reduced over time along with an increase in PARP cleavage. [3] A recent research indicates NX-2112 induces autophagy in a time- and dose-dependent manner via Akt/mTOR/p70S6K inhibition. SNX-2112 induces significant apoptosis and utophagy in human melanoma A-375 cells, and may be an effective targeted therapy agent. [4] |
| 体内活性 | SNX-2112, delivered by its prodrug SNX-5422, inhibits MM cell growth and prolongs survival in a xenograft murine model and blockade of Hsp90 by SNX-2112 not only inhibits MM cell growth but also acts in the bone marrow microenvironment to block angiogenesis and osteoclastogenesis. [2] |
| 激酶实验 | ATP Displacement Assay: For the protein affinity-displacement assay, a purine-based affinity resin is generated by incubating ATP-linked Sepharose with Jurkat cell lysate (flash frozen and homogenized in saline) at 4 °C. This is then incubated with SNX-2112 for 90 minutes. Proteins eluted by drug are then resolved by SDS-PAGE, visualized with silver staining, and excised from the gel for MS-based identification. Briefly, after destaining and trypsin digestion, peptides are extracted with μC18 ZipTips and then eluted and spotted directly to a conventional stainless steel matrix-assisted laser desorption/ionization target with a saturated solution of α-cyano-4- hydroxycinnamic acid in 50% acetonitrile, 0.15% formic acid. Mass spectra are then acquired using a MALDI-TOF/TOF 4700 Proteomics Analyzer. MS spectra are acquired (1,000 shots per spectrum), and the three peaks from each with the greatest signal-to-noise ratio are automatically submitted for tandem MS analysis (3000 shots per spectrum). The collision energy is 1keV. Air is used as the collision gas. Protein identification is done from the MS and tandem MS data using GPS Explorer software with the integrated Mascot database search engine. |
| 细胞实验 | Cell viability is determined by seeding 2-5 × 103 cells per well in 96- well plates and treating with SNX-2112 24 hours after plating in complete medium (200 μL). Each drug concentration is tested in eight wells. Cells are assayed using the Alamar blue viability test after a 96-h incubation. Flow cytometry is done using nuclei stained with ethidium bromide and isolated via the Nusse protocol(Only for Reference) |
| 别名 | SNX-2112, PF 04928473, SNX 2112 |
| 分子量 | 464.48 |
| 分子式 | C23H27F3N4O3 |
| CAS No. | 908112-43-6 |
Powder: -20°C for 3 years | In solvent: -80°C for 1 year
Ethanol: 1 mg/mL (2.15 mM)
DMSO: 86 mg/mL (185.2 mM)
H2O: < 1 mg/mL (insoluble or slightly soluble)
| 可选溶剂 | 浓度 体积 质量 | 1 mg | 5 mg | 10 mg | 25 mg |
| Ethanol / DMSO | 1 mM | 2.1529 mL | 10.7647 mL | 21.5295 mL | 53.8236 mL |
| DMSO | 5 mM | 0.4306 mL | 2.1529 mL | 4.3059 mL | 10.7647 mL |
| 10 mM | 0.2153 mL | 1.0765 mL | 2.1529 mL | 5.3824 mL | |
| 20 mM | 0.1076 mL | 0.5382 mL | 1.0765 mL | 2.6912 mL | |
| 50 mM | 0.0431 mL | 0.2153 mL | 0.4306 mL | 1.0765 mL | |
| 100 mM | 0.0215 mL | 0.1076 mL | 0.2153 mL | 0.5382 mL |
对于不同动物的给药剂量换算,您也可以参考 更多...
请在以下方框中输入您的动物实验信息后点击计算,可以得到母液配置方法和体内配方的制备方法: 比如您的给药剂量是10 mg/kg,每只动物体重20 g,给药体积100 μL,一共给药动物10 只,您使用的配方为5% DMSO+30% PEG300+5% Tween 80+60% ddH2O。那么您的工作液浓度为2 mg/mL。
母液配置方法:2 mg 药物溶于 50 μL DMSO (母液浓度为 40 mg/mL), 如您需要配置的浓度超过该产品的溶解度,请先与我们联系。
体内配方的制备方法:取 50 μL DMSO 主液,加入 300 μL PEG300, 混匀澄清,再加 50 μL Tween 80,混匀澄清,再加 600 μL ddH2O, 混匀澄清。
您可能有的问题的答案可以在抑制剂处理说明中找到,包括如何准备库存溶液,如何存储产品,以及基于细胞的分析和动物实验需要特别注意的问题。
SNX2112 908112-43-6 Cytoskeletal Signaling Metabolism HSP PF04928473 SNX-2112 PF-04928473 PF 04928473 SNX 2112 Inhibitor inhibitor inhibit
