Powder: -20°C for 3 years | In solvent: -80°C for 1 year
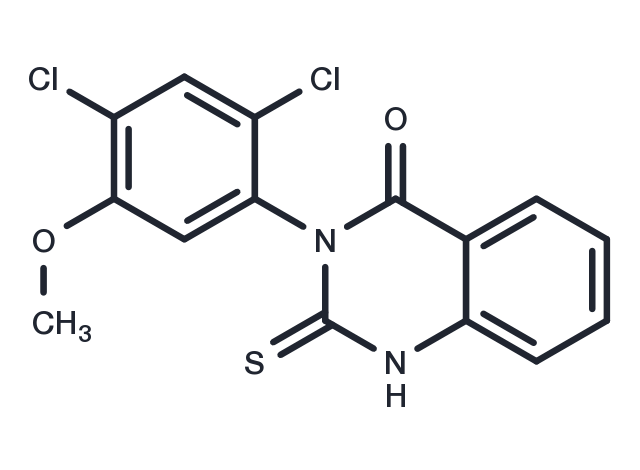
Mdivi-1 (Mitochondrial division inhibitor 1) 是一种线粒体分裂抑制剂,抑制 DRP1 和 Dynamin I (IC50=1-10 μM)。Mdivi-1 可以抑制线粒体自噬。
| 规格 | 价格/CNY | 货期 | 数量 | |
|---|---|---|---|---|
| 1 mg | ¥ 153 | 现货 | ||
| 5 mg | ¥ 315 | 现货 | ||
| 10 mg | ¥ 523 | 现货 | ||
| 25 mg | ¥ 987 | 现货 | ||
| 50 mg | ¥ 1,790 | 现货 | ||
| 100 mg | ¥ 2,530 | 现货 | ||
| 200 mg | ¥ 3,730 | 现货 | ||
| 500 mg | ¥ 5,950 | 现货 | ||
| 1 mL * 10 mM (in DMSO) | ¥ 315 | 现货 | ||
| 产品描述 | Mdivi-1 (Mitochondrial division inhibitor 1) is a mitochondrial division inhibitor that inhibits DRP1 and Dynamin I (IC50=1-10 μM). Mdivi-1 inhibits mitochondrial autophagy. |
| 靶点活性 | Dnm1 GTPase:1 μM-10 μM |
| 体外活性 |
方法:人乳腺癌细胞 MDA-MB-231 用 Mdivi-1 (5-40 μM) 处理 8 h,使用 Immunostaining 方法分析有丝分裂纺锤体的异常。 结果:Mdivi-1 剂量依赖性地增加具有纺锤体异常的有丝分裂细胞的百分比。[1] 方法:人肺癌细胞 H460、A549 和人结直肠癌细胞 HCT116 用 Mdivi-1 (20-50 μM) 处理 24 h,使用 Flow Cytometry 方法检测细胞死亡情况。 结果:Mdivi-1 显著抑制三种肿瘤细胞的死亡。[2] |
| 体内活性 |
方法:为检测体内活性,将 Mdivi-1 (50 mg/kg) 腹腔注射给 C57BL/6 小鼠,随后通过急性眼压升高诱导短暂的视网膜缺血。 结果:Mdivi-1 治疗阻断了缺血视网膜中的凋亡细胞死亡,并显著提高了缺血后两周视网膜神经节细胞的存活率。[3] 方法:为研究对实验性自身免疫性脑脊髓炎 (EAE) 的影响,将 Mdivi-1 (25 mg/kg) 腹腔注射给诱导 EAE 的 C57BL/6 小鼠,每天一次,持续二十五天。 结果:Mdivi-1 通过减少中枢神经系统中的脱髓鞘和细胞浸润,有效地抑制了 EAE 的严重程度。Mdivi-1 通过调节 Th1/Th17 和调节性 T 细胞之间的平衡,在 EAE 中具有治疗潜力。[4] |
| 激酶实验 | All GTPase assay reactions are started in a 200 μL volume, of which 150 μL is placed into the well of a 96-well plate. Depletion of NADH, as monitored by reading the A340 of the reaction, is measured every 20 s for a total of 40 min using a SpectraMAX 250 96-well plate reader. Spectrophotometric data are transferred to Excel and the measured steady state depletion of NADH over time is converted to protein activity. |
| 细胞实验 | Mdivi-1 is dissolved in DMSO. YPGlycerol plates are topped with 10 mL YPGlycerol containing 1% low melt agar and 75 μM mdivi-1, and cells are spotted 12 hours later using a 48 well pinning device. After pinning cells, plates are incubated at 24°C or 37°C and imaged using an Eagle Eye II imaging system. |
| 别名 | Mitochondrial division inhibitor 1 |
| 分子量 | 353.22 |
| 分子式 | C15H10Cl2N2O2S |
| CAS No. | 338967-87-6 |
Powder: -20°C for 3 years | In solvent: -80°C for 1 year
DMSO: 35.3 mg/mL (100 mM)
| 可选溶剂 | 浓度 体积 质量 | 1 mg | 5 mg | 10 mg | 25 mg |
| DMSO | 1 mM | 2.8311 mL | 14.1555 mL | 28.311 mL | 70.7774 mL |
| 5 mM | 0.5662 mL | 2.8311 mL | 5.6622 mL | 14.1555 mL | |
| 10 mM | 0.2831 mL | 1.4155 mL | 2.8311 mL | 7.0777 mL | |
| 20 mM | 0.1416 mL | 0.7078 mL | 1.4155 mL | 3.5389 mL | |
| 50 mM | 0.0566 mL | 0.2831 mL | 0.5662 mL | 1.4155 mL | |
| 100 mM | 0.0283 mL | 0.1416 mL | 0.2831 mL | 0.7078 mL |
对于不同动物的给药剂量换算,您也可以参考 更多...
请在以下方框中输入您的动物实验信息后点击计算,可以得到母液配置方法和体内配方的制备方法: 比如您的给药剂量是10 mg/kg,每只动物体重20 g,给药体积100 μL,一共给药动物10 只,您使用的配方为5% DMSO+30% PEG300+5% Tween 80+60% ddH2O。那么您的工作液浓度为2 mg/mL。
母液配置方法:2 mg 药物溶于 50 μL DMSO (母液浓度为 40 mg/mL), 如您需要配置的浓度超过该产品的溶解度,请先与我们联系。
体内配方的制备方法:取 50 μL DMSO 主液,加入 300 μL PEG300, 混匀澄清,再加 50 μL Tween 80,混匀澄清,再加 600 μL ddH2O, 混匀澄清。
您可能有的问题的答案可以在抑制剂处理说明中找到,包括如何准备库存溶液,如何存储产品,以及基于细胞的分析和动物实验需要特别注意的问题。
Mdivi-1 338967-87-6 Apoptosis Autophagy Cytoskeletal Signaling Mitophagy Dynamin Mitochondrial division inhibitor 1 Mitochondrial Autophagy Mitochondrial division inhibitor1 Inhibitor inhibit Mdivi 1 Mdivi1 Mitochondrial division inhibitor-1 inhibitor
